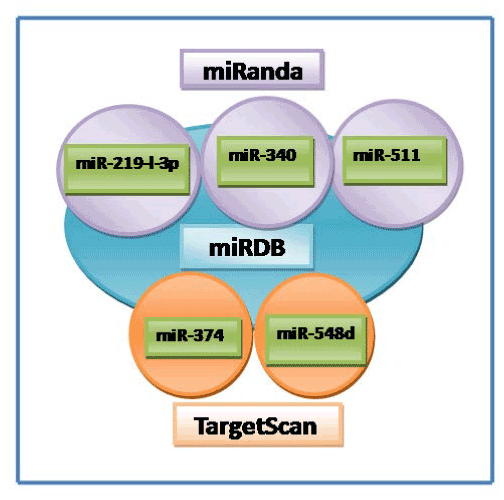
Computational prediction of novel human miRNAs/miRNA target sites in correlation with SNP (rs678653) at 3'UTR of cyclin D1 gene

Prediction methods for microRNA targets in bilaterian animals: Toward a better understanding by biologists - ScienceDirect

MIPDH: A Novel Computational Model for Predicting microRNA–mRNA Interactions by DeepWalk on a Heterogeneous Network | ACS Omega

Target prediction and validation of microRNAs expressed from FSHR and aromatase genes in human ovarian granulosa cells | Scientific Reports

Too Many False Targets for MicroRNAs: Challenges and Pitfalls in Prediction of miRNA Targets and Their Gene Ontology in Model and Non‐model Organisms - Fridrich - 2019 - BioEssays - Wiley Online Library

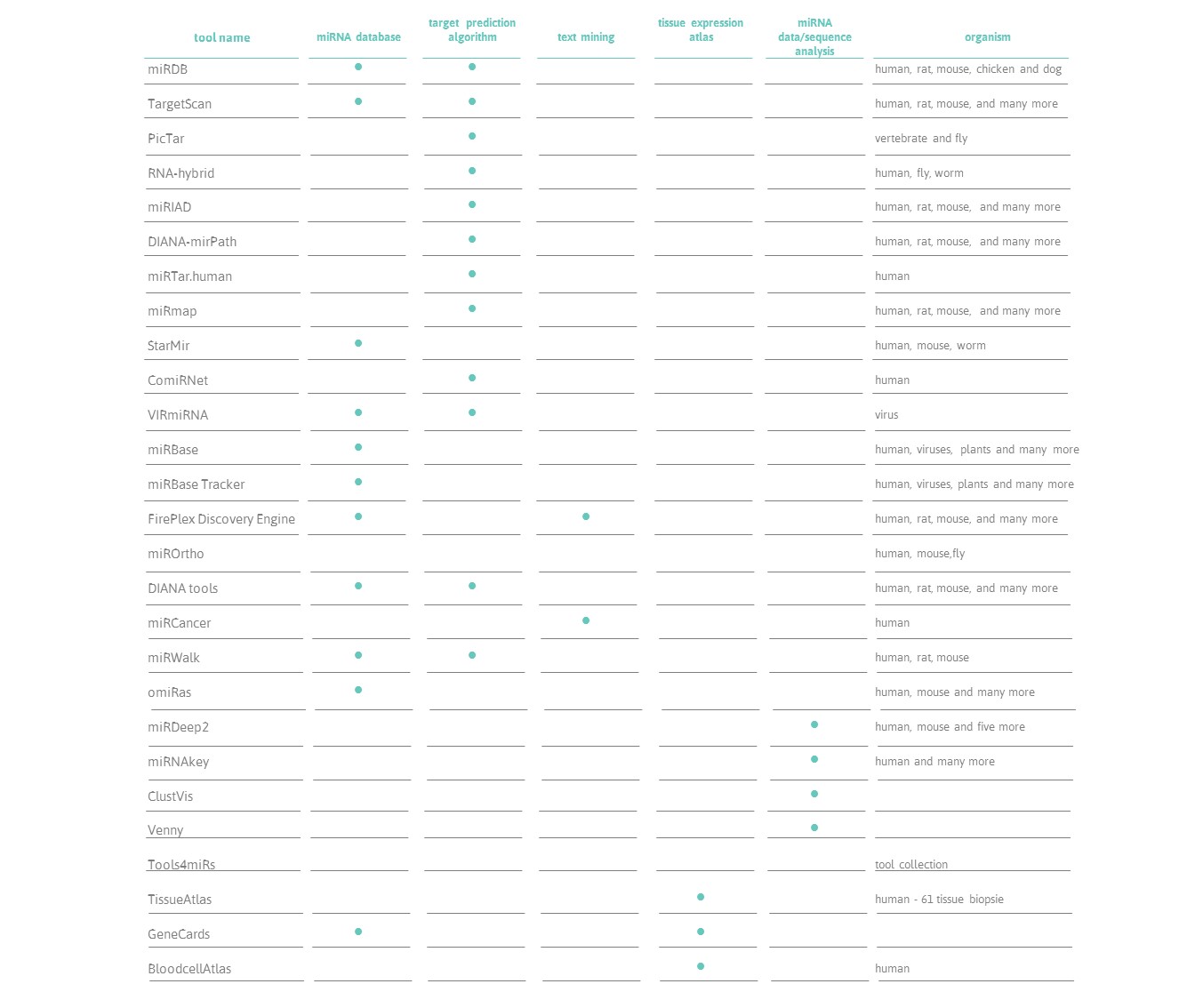
Popular Computational Tools Used for miRNA Prediction and Their Future Development Prospects | SpringerLink

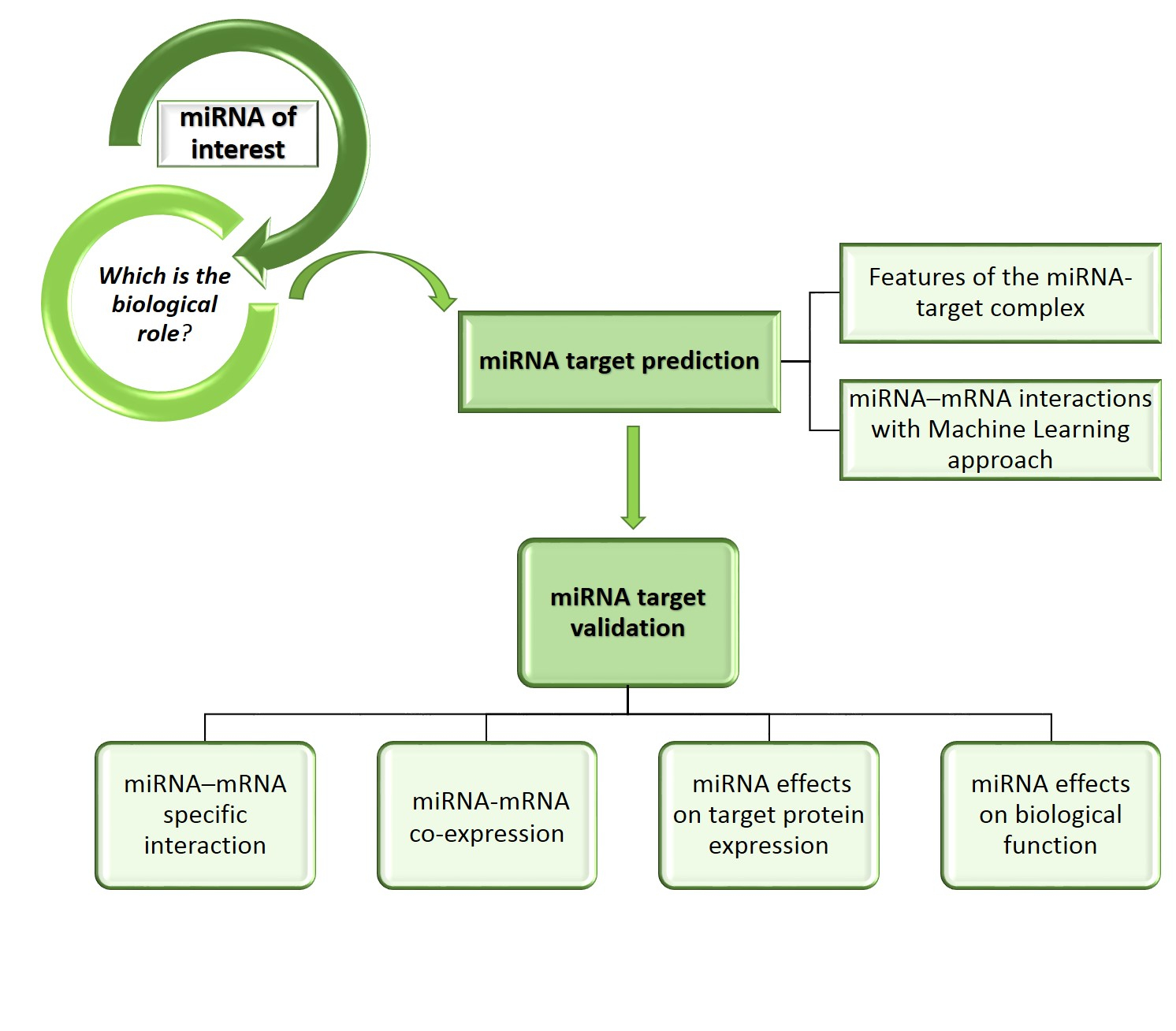
miRNA Target Prediction and Integration Workflow. The miRNA Targets... | Download Scientific Diagram

RWRMTN: a tool for predicting disease-associated microRNAs based on a microRNA-target gene network | BMC Bioinformatics | Full Text

Genome wide in-silico miRNA and target network prediction from stress responsive Horsegram (Macrotyloma uniflorum) accessions | Scientific Reports
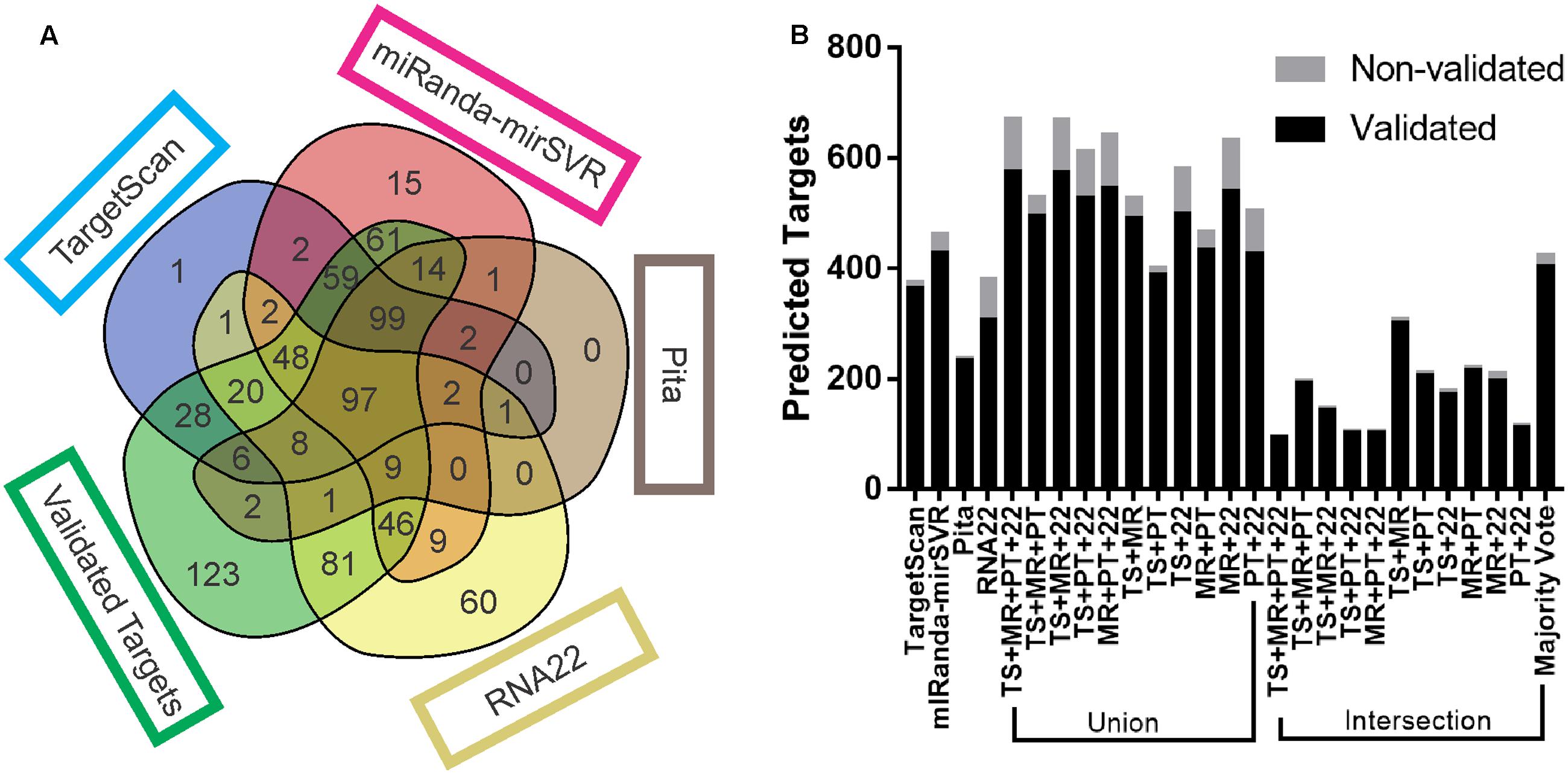
Frontiers | Combining Results from Distinct MicroRNA Target Prediction Tools Enhances the Performance of Analyses